Abstract
Introduction: This review aims to present briefly the new horizon opened to pharmacology by the deep learning (DL) technology, but also to underline the most important threats and limitations of this method.
Material and Methods: We searched multiple databases for articles published before May 2021 according to the preferred reported item related to deep learning and drug research. Out of the 267 articles retrieved, we included 49 in the final review.
Results: DL and other different types of artificial intelligence have recently entered all spheres of science, taking an increasingly central position in the decision-making processes, also in pharmacology. Hence, there is a need for better understanding of these technologies. The basic differences between AI (artificial intelligence), DL and ML (machine learning) are explained. Additionally, the authors try to highlight the role of deep learning methods in drug research and development as well as in improving the safety of pharmacotherapy. Finally, future directions of DL in pharmacology were outlined as well as possible misuses of it.
Conclusion: DL is a promising and powerful tool for comprehensive analysis of big data related to all fields of pharmacology, however it has to be used carefully.
Citation
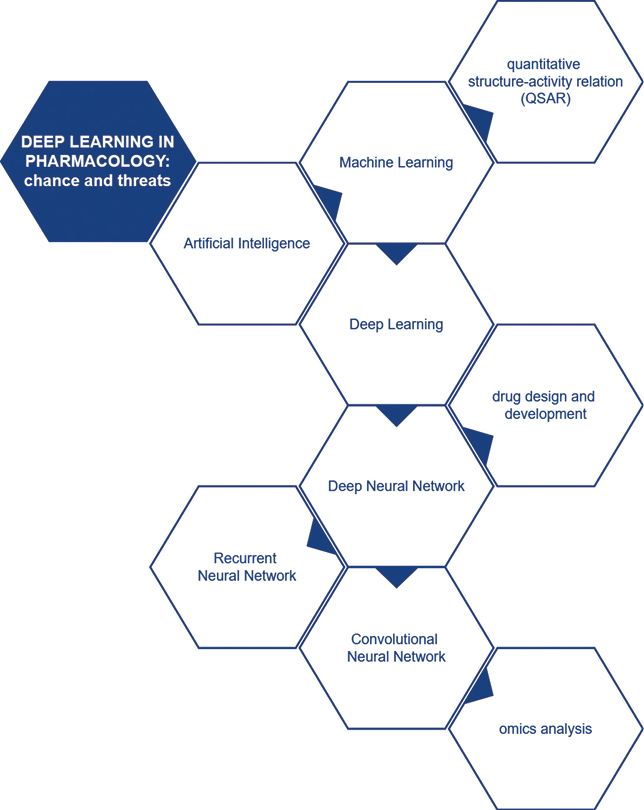
Kocić I, Kocić M, Rusiecka I, Kocić A, Kocić E. Deep learning in pharmacology: opportunities and threats. Eur J Transl Clin Med. 2022;5(2):88-94Figure 1. Summary graph
Introduction
Despite a widely accepted opinion that artificial intelligence (AI) era started with Alan Turing’s publication of “Computing Machinery and Intelligence” in 1950 [1], discussion about that kind of human vision described with mathematical symbols appeared for the first time in the XVII century with works by Leibniz, Hobbes and Descartes. However, the practical implementation of AI into daily life is fairly new, as it began in the 1990s [2]. There are several popular terms used to describe AI-related new technologies entering recently in almost all spheres of sciences and everyday life: artificial intelligence, machine learning (ML) and deep learning (DL) [3-4]. Of the three, AI is the broadest concept, encompassing both ML and DL. The basic characteristic of all of these new technologies is big data management, which allows to find out specific correlation patterns, invisible for simple algorithms, and statistical evaluation with limited data. ML uses algorithmic models, which treat the data as an unknown and find generalizable predictive patterns, while statistical modeling assumes that the data is generated by a given random data model and draws population inferences from a sample [5-6].
On the other hand, DL, a subset of ML, structures algorithms in layers to create deep neural network with many hidden layers, which provide better pattern recognition and new possibilities in data mining [7-9]. Specifically, the aim of DL is to determine a mathematical function f that maps a number of inputs (x) to their corresponding outputs (y), e.g. y = f(x). In other words, standard network architecture of neural networks contains an input layer, several hidden layers in between and an output layer. A set of training data, often called “batch” is fed forward through the network’s layers and the output layer computes the loss function as the difference between the calculated prediction and the correct response. After that, loss error of the next operation is reduced and a backpropagation algorithm adjusts filter banks and learns the value of the parameter resulting in the best function approximation [9]. Recurrent neural networks, derived from feedforward networks, do not use limited size of context, which allows information to cycle as long as needed. This makes them useful in sequential data prediction such as language modelling [10].
Another network type is the convolutional neural network (CNN), which is widely used in systems that deal with image classification and computer vision in general. CNNs are composed of multiple types of hidden layers: convolutional layers, pooling layers and fully-connected layers. Like in the other network types, the input layer contains the input data, i.e. the pixel values of the image. The convolutional layer calculates the scalar product between the weights and the region connected to the input volume. The rectified linear unit (ReLU) applies an “elementwise” activation function such as sigmoid to previous layer’s activation output. The pooling layer performs downsampling along input’s spatial dimensionality, reducing the number of parameters within activation. The fully-connected layers will then attempt to produce scores for classification from the activations. This is only the base architecture model – as CNNs often deal with very complex image data, optimisations are often necessary [11]. Some new deep learning approaches incorporate fusion strategies into the deep learning architecture itself, creating fuzzy hidden layers. These layers are able to condense hundreds of inputs into a more manageable set. Fusion can offer major reduction in model complexity [12].
As Schmidhuber et al. suggested, the primary deficiency of most traditional ML methods as compared to DL methods, is that they have a limited ability to simulate a complicated approximation function and generalize to an unseen instance [9]. Usually, ML is used for supervised analysis and DL for more complex, unsupervised ones. Matter of fact, DL methods can be used both in supervised applications- for accurate prediction of one or more labels or outcomes associated with each data point (instead of a simple regression approaches), as well as in unsupervised (or ‘exploratory’) applications, where the goal is to summarize, explain or identify appropriate patterns in a dataset as a form of clustering. Moreover, DL methods may combine both of these approaches and propose feature oriented one, as a highly precise predictor [13].
ML and DL are widely used in daily life, e.g. to improve street traffic safety, in marketing research and, what is seen as a very controversial issue, to determine the voters’ preferences [14]. Medicine as a science and clinical discipline is not an exception. DL method is a very promising tool in the diagnostic procedures (e.g. in pathomorphology and X-ray imaging) and results interpretation, but needs further research. For instance, Wang et al. analysed stained slides of lymph node slices to identify cancers, and found out that a pathologist had an error rate of about 3%, while the applied algorithm had about 6%. The pathologist did not produce any false positives but did have a number of false negatives. The algorithm had about twice the error rate of a pathologist, but the errors were not strongly correlated [14]. Academic institutions and start-ups alike are rapidly developing prototype technologies using the data of healthcare providers, individuals, and healthcare organizations’, however the ethical implications, vulnerabilities and potential for misuse of such tools are still not taken seriously [15].
The value of algorithms proposed by DL methods is directly dependent on the quality and quantity of the entry data. It seems that a DL analytical platform which has got thousands of microscopic samples or X-ray images of lung changes during pneumonia or kidney cancer will unmistakably recognize the next one. However, recent controversies with face recognition technology (accurate only for faces of white men, whereas 20- 30% recognition of Asian women’s faces) raise new questions about practical usage of image recognition technologies [16]. Currently, these new technologies are present in almost all areas of medicine, including pharmacology. This review aims to present briefly the new horizon opened to pharmacology by the deep learning (DL) technology, but also to underline the most important threats and limitations of this method.
Material and methods
The PubMed, EMBASE and Cochrane Library were searched for articles published before May 2021 according to the preferred reported item related to deep learning and drug research. The following keywords were applied: artificial intelligence, deep learning, machine learning and drug research, research and development. Articles were included in the analysis based on their quality and journal rank.
Results
Out of the 267 articles retrieved, we included 49 in the final review. No statistical analysis was performed.
Discussion
DL in pharmacology
There are many different aspects of possible use of DL as a tool able to create predictive models and recognize complex patterns in big data sets in drug research and development. A serious challenge for DL technology is how to manage the huge amounts of data obtained by omics technologies (e.g. metabolomics, genomics, proteomics, glycomics) in order to support and improve personal approach of pharmacotherapy. In this context, an up to date DL method is successfully used in sequence analysis, genome wide association studies, transcriptomics, epigenomics, proteomics and metabolomics. It seems that the convolutional neural network (CNN) DL model is the most suitable for omics analysis as a tool with a transfer learning strategy (transferring prior knowledge from a source domain into a target domain) dealing with relatively small data sets. Using this, it is possible to detect single nucleotide polymorphism (SNP) to predict the relation between genetic variants and gene expression, as well as to predict regulatory motifs in the genome and promoter sequences in the gene [17-22].
Pharmacology as a multidisciplinary science is practically involved in every clinical discipline of medicine (e.g. in surgery via the use of analgesics, anesthetics and antibiotics), hence special attention should be paid to it. Drug discovery and development is a complex process involving many different techniques. Traditional ML methods have already been widely used by pharmacologists in quantitative structure-activity relation (QSAR) models (Fig. 2). However, DL algorithms in drug design and development are slowly becoming dominant due to improved feasibility of computational management of the enormous amounts of chemical data involved [23-28].
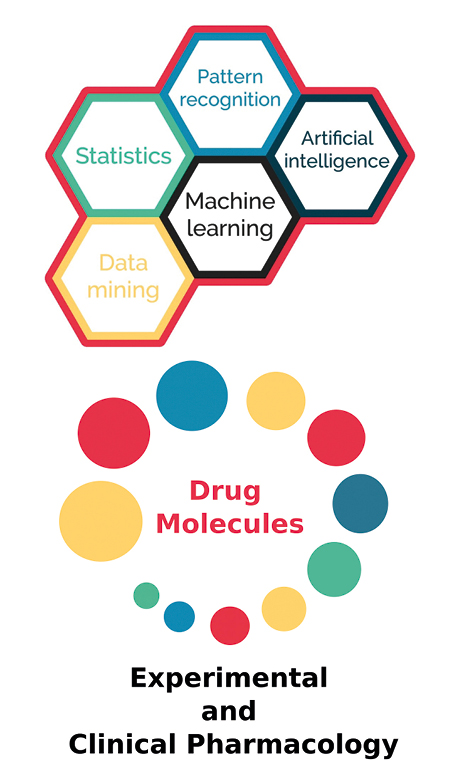

Figure 2. Deep learning in pharmacology
Generally, there are three types of DL networks used in drug discovery: CNN, RNN and DNN. So far, many DL models for preclinical research have been reported. They predict the drug-target interaction of novel drug molecules (drug design) with theirs absorption, distribution, metabolism, excretion, and toxicity (ADMET) [29]. Interesting DL models have been developed for drug discovery requiring 3D structure for both ligand and target, known as AtomNet [30]. Thanks to them it is possible to predict binding affinity for a selective active compound. The next step of using DL models in drug discovery process was modelling of drug mechanism of action (MOA) using genetic profile activity relationship (GPAR) [31]. In order to generate more reliable MOA hypothesis, one can use GPAR to customize its training set to train MOA prediction models and to evaluate the model performances. An interesting example of using DL for prediction of the new drug molecules to be further tested in industrial environment was published by Sturm et al [32]. They found that evaluation of ExCAPE-DB, one of the largest open-source benchmark target prediction datasets, allows to evaluate target prediction models trained on public data and predict industrial QSAR, therefore it is usable in industrial drug discovery projects.
A major challenge is an attempt to use AI and DL in order to improve the safety of pharmacotherapy. A part from posing serious health hazards and generating enormous financial costs, dverse drug reactions (ADRs) are preventable. There is a hope that DL can help in much better and effective prediction and significant reduction of ADR [33]. Finally, optimal medication dosing can be also predicted by using appropriate, publicly available big data and deep reinforcement learning approach [34]. Prediction of optimal and safe treatment of patients is a completely new field for DL application. Nevertheless, new methods to recommend patient treatment and predict its outcome as well as to identify drug targets and predict drug response and interactions are being developed [35-37]. Another prospective in development of DL in pharmacology is personalization of pharmacotherapy. For instance, the genetic expression profile-based screening of appropriate therapeutics for cancer treatment sis already in development (OncoFinder algorithm) [38].
Conclusions
False interpretation of obtained data and misunderstanding of DL technology are the real and serious threats of using of DL as an advisor for personalizing of pharmacotherapy in context of type of drug and optimal dosing. Some of the most important pros and cons of DL are summarized in Table 1. Moreover, in basic biological research, measures of uncertainty help researchers distinguish between true regularities in the data and patterns that are false or uncertain. Two main types of uncertainties used in calculations are epistemic and aleatoric uncertainties [18]. Epistemic uncertainty is a measure of uncertainty concerning the model, including its structure and parameters. It is caused by the lack of sufficient training data, so that it can be reduced with better access to more data with better quality. In contrast, aleatoric uncertainty is a description of uncertainty of observations, due to the noise or missing parts in data. It diminishes with improvements in the measurement precision of the data [39].
Progress in decision making using deep learning is often set back by the fact that predicting results of a set of circumstances is much simpler than that based on a desirable outcome. There still remain many problems such as low interpretability of models and dealing with limited and mixed data in dynamic settings. Interpretability problems are particularly important as decision making in medicine is understandably averse to risk, and with low interpretability it is difficult to reason about the model and build trust in its correctness. While ML models also present some dangerous vulnerabilities, like misclassification of adversarial examples [40-41], there are constant improvements in this area [42-44]. Cooperation between human experts and DLbased systems seems to offer the best results for alleviating many problems [45-48]. Obviously, ethical issues are also extremely important in this context and can seriously restrict the use and publishing of very sensitive health related data [49].
Funding
This work was supported by the Faculty of Medicine of the Medical University of Gdańsk (ST-02-00-22/07).
Conflicts of interest
None.
References
| 1. |
Turing AM. Computing machinery and intelligence Mind 49 433-460. Mind. 1950;59(236):433–60.
|
| 2. |
Clark J. Why 2015 Was a Breakthrough Year in Artificial Intelligence [Internet]. Bloomberg. Europe Edition. 2015 [cited 2022 Aug 22]. Available from: https://www.bloomberg.com/news/articles/2015-12-08/why-2015-was-a-breakthrough-year-in-artificial-intelligence.
|
| 3. |
Cukier K. Ready for robots? How to think about the future of AI [Internet]. Foreign Affairs. [cited 2022 Aug 22]. Available from: https://www.foreignaffairs.com/reviews/review-essay/2019-06-11/ready-robots.
|
| 4. |
McCorduck P, Cfe C. Machines Who Think: A Personal Inquiry into the History and Prospects of Artificial Intelligence [Internet]. CRC Press; 2004. Available from: https://books.google.pl/books?id=r2C1DwAAQBAJ.
|
| 5. |
Jang H, Park A, Jung K. Neural Network Implementation Using CUDA and OpenMP. In: 2008 Digital Image Computing: Techniques and Applications [Internet]. IEEE; 2008. p. 155–61. Available from: https://doi.org/10.1109/DICTA.2008.82.
|
| 6. |
Bzdok D, Altman N, Krzywinski M. Statistics versus machine learning. Nat Methods [Internet]. 2018 Apr 3;15(4):233–4. Available from: https://doi.org/10.1038/nmeth.4642.
|
| 7. |
Hollis KF, Soualmia LF, Séroussi B. Artificial Intelligence in Health Informatics: Hype or Reality? Yearb Med Inform [Internet]. 2019 Aug 16;28(01):003–4. Available from: https://doi.org/10.1055/s-0039-1677951.
|
| 8. |
Lavecchia A. Machine-learning approaches in drug discovery: methods and applications. Drug Discov Today [Internet]. 2015 Mar;20(3):318–31. Available from: https://www.sciencedirect.com/science/article/pii/S1359644614004176.
|
| 9. |
Schmidhuber J. Deep learning in neural networks: An overview. Neural Networks [Internet]. 2015;61:85–117. Available from: https://www.sciencedirect.com/science/article/pii/S0893608014002135.
|
| 10. |
Mikolov T. Recurrent Neural Network based Language Model research paper [Internet]. 2010. Available from: http://www.fit.vutbr.cz/research/groups/speech/servite/2010/rnnlm_mikolov.pdf.
|
| 11. |
O’Shea K, Nash R. An Introduction to Convolutional Neural Networks [Internet]. Cornell University. 2015 [cited 2023 Aug 22]. Available from: https://arxiv.org/abs/1511.08458.
|
| 12. |
Price SR, Price SR, Anderson DT. Introducing Fuzzy Layers for Deep Learning. In: 2019 IEEE International Conference on Fuzzy Systems (FUZZ-IEEE) [Internet]. IEEE; 2019. p. 1–6. Available from: https://ieeexplore.ieee.org/document/8858790/.
|
| 13. |
Cao C, Liu F, Tan H, Song D, Shu W, Li W, et al. Deep Learning and Its Applications in Biomedicine. Genomics Proteomics.
|
| 14. |
Bioinformatics [Internet]. 2018;16(1):17–32. Available from: https://www.sciencedirect.com/science/article/pii/S1672022918300020.
|
| 15. |
Doucette J, Larson K, Cohen R. Conventional Machine Learning for Social Choice. Proc AAAI Conf Artif Intell [Internet]. 2015 Feb 16;29(1 SE-AAAI Technical Track: Game Theory and Economic Paradigms). Available from: https://ojs.aaai.org/index.php/AAAI/article/view/9294.
|
| 16. |
Papernot N, McDaniel P, Sinha A, Wellman MP. SoK: Security and Privacy in Machine Learning. In: 2018 IEEE European Symposium on Security and Privacy (EuroS&P) [Internet]. IEEE; 2018. p. 399–414. Available from: https://ieeexplore.ieee.org/document/8406613/.
|
| 17. |
Zou J, Schiebinger L. AI can be sexist and racist — it’s time to make it fair. Nature [Internet]. 2018 Jul 18;559(7714):324–6. Available from: http://www.nature.com/articles/d41586-018-05707-8.
|
| 18. |
Zhang L, Tan J, Han D, Zhu H. From machine learning to deep learning: progress in machine intelligence for rational drug discovery. Drug Discov Today [Internet]. 2017 Nov;22(11):1680–5. Available from: https://linkinghub.elsevier.com/retrieve/pii/S1359644616304366.
|
| 19. |
Ching T, Himmelstein DS, Beaulieu-Jones BK, Kalinin AA, Do BT, Way GP, et al. Opportunities and obstacles for deep learning in biology and medicine. J R Soc Interface [Internet]. 2018 Apr 4;15(141):20170387. Available from: https://royalsocietypublishing.org/doi/10.1098/rsif.2017.0387.
|
| 20. |
Wang D, Khosla A, Gargeya R, Irshad H, Beck AH. Deep learning for identifying metastatic breast cancer. Cornell Univ [Internet]. 2016; Available from: https://arxiv.org/abs/1606.05718.
|
| 21. |
Arnedo J, del Val C, de Erausquin GA, Romero-Zaliz R, Svrakic D, Cloninger CR, et al. PGMRA: a web server for (phenotype x genotype) many-to-many relation analysis in GWAS. Nucleic Acids Res [Internet]. 2013 Jul 1;41(W1):W142–9. Available from: https://academic.oup.com/nar/article-lookup/doi/10.1093/nar/gkt496.
|
| 22. |
Hill ST, Kuintzle R, Teegarden A, Merrill E, Danaee P, Hendrix DA. A deep recurrent neural network discovers complex biological rules to decipher RNA protein-coding potential. Nucleic Acids Res [Internet]. 2018 Sep 19;46(16):8105–13. Available from: https://academic.oup.com/nar/article/46/16/8105/5050624.
|
| 23. |
Tripathi R, Patel S, Kumari V, Chakraborty P, Varadwaj PK. DeepLNC, a long non-coding RNA prediction tool using deep neural network. Netw Model Anal Heal Informatics Bioinforma [Internet]. 2016 Dec 10;5(1):21. Available from: http://link.springer.com/10.1007/s13721-016-0129-2.
|
| 24. |
Alipanahi B, Delong A, Weirauch MT, Frey BJ. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat Biotechnol [Internet]. 2015 Aug 27;33(8):831–8. Available from: http://www.nature.com/articles/nbt.3300.
|
| 25. |
Date Y, Kikuchi J. Application of a Deep Neural Network to Metabolomics Studies and Its Performance in Determining Important Variables. Anal Chem [Internet]. 2018 Feb 6;90(3):1805–10. Available from: https://pubs.acs.org/doi/10.1021/acs.analchem.7b03795.
|
| 26. |
Asakura T, Date Y, Kikuchi J. Application of ensemble deep neural network to metabolomics studies. Anal Chim Acta [Internet]. 2018 Dec;1037:230–6. Available from: https://linkinghub.elsevier.com/retrieve/pii/S0003267018302605.
|
| 27. |
Svetnik V, Liaw A, Tong C, Culberson JC, Sheridan RP, Feuston BP. Random Forest: A Classification and Regression Tool for Compound Classification and QSAR Modeling. J Chem Inf Comput Sci [Internet]. 2003 Nov 1;43(6):1947–58. Available from: https://pubs.acs.org/doi/10.1021/ci034160g.
|
| 28. |
Goh GB, Hodas NO, Vishnu A. Deep learning for computational chemistry. J Comput Chem [Internet]. 2017 Jun 15;38(16):1291–307. Available from: https://onlinelibrary.wiley.com/doi/10.1002/jcc.24764.
|
| 29. |
Chen B, Sheridan RP, Hornak V, Voigt JH. Comparison of Random Forest and Pipeline Pilot Naïve Bayes in Prospective QSAR Predictions. J Chem Inf Model [Internet]. 2012 Mar 26;52(3):792–803. Available from: https://pubs.acs.org/doi/10.1021/ci200615h.
|
| 30. |
Myint KZ, Xie X-Q. Ligand Biological Activity Predictions Using Fingerprint-Based Artificial Neural Networks (FANN-QSAR). In 2015. p. 149–64. Available from: http://link.springer.com/10.1007/978-1-4939-2239-0_9.
|
| 31. |
Ma C, Wang L, Yang P, Myint KZ, Xie X-Q. LiCABEDS II. Modeling of Ligand Selectivity for G-Protein-Coupled Cannabinoid Receptors. J Chem Inf Model [Internet]. 2013 Jan 28;53(1):11–26. Available from: https://pubs.acs.org/doi/10.1021/ci3003914.
|
| 32. |
Gray KA, Yates B, Seal RL, Wright MW, Bruford EA. Genenames.org: the HGNC resources in 2015. Nucleic Acids Res [Internet]. 2015 Jan 28;43(D1):D1079–85. Available from: http://academic.oup.com/nar/article/43/D1/D1079/2437320/Genenamesorg-the-HGNC-resources-in-2015.
|
| 33. |
Sturm N, Mayr A, Le Van T, Chupakhin V, Ceulemans H, Wegner J, et al. Industry-scale application and evaluation of deep learning for drug target prediction. J Cheminform [Internet]. 2020 Dec 19;12(1):26. Available from: https://jcheminf.biomedcentral.com/articles/10.1186/s13321-020-00428-5.
|
| 34. |
Basile AO, Yahi A, Tatonetti NP. Artificial Intelligence for Drug Toxicity and Safety. Trends Pharmacol Sci [Internet]. 2019 Sep;40(9):624–35. Available from: https://linkinghub.elsevier.com/retrieve/pii/S0165614719301427.
|
| 35. |
Nemati S, Ghassemi MM, Clifford GD. Optimal medication dosing from suboptimal clinical examples: A deep reinforcement learning approach. In: 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) [Internet]. IEEE; 2016. p. 2978–81. Available from: http://ieeexplore.ieee.org/document/7591355/.
|
| 36. |
Ferreira LLG, Andricopulo AD. ADMET modeling approaches in drug discovery. Drug Discov Today [Internet]. 2019 May;24(5):1157–65. Available from: https://linkinghub.elsevier.com/retrieve/pii/S1359644618303301.
|
| 37. |
Tsou LK, Yeh S-H, Ueng S-H, Chang C-P, Song J-S, Wu M-H, et al. Comparative study between deep learning and QSAR classifications for TNBC inhibitors and novel GPCR agonist discovery. Sci Rep [Internet]. 2020 Dec 8;10(1):16771. Available from: https://www.nature.com/articles/s41598-020-73681-1.
|
| 38. |
Gao S, Han L, Luo D, Liu G, Xiao Z, Shan G, et al. Modeling drug mechanism of action with large scale gene-expression profiles using GPAR, an artificial intelligence platform. BMC Bioinformatics [Internet]. 2021 Dec 7;22(1):17. Available from: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-020-03915-6.
|
| 39. |
Aliper A, Plis S, Artemov A, Ulloa A, Mamoshina P, Zhavoronkov A. Deep Learning Applications for Predicting Pharmacological Properties of Drugs and Drug Repurposing Using Transcriptomic Data. Mol Pharm [Internet]. 2016 Jul 5;13(7):2524–30. Available from: https://pubs.acs.org/doi/10.1021/acs.molpharmaceut.6b00248.
|
| 40. |
Wen M, Zhang Z, Niu S, Sha H, Yang R, Yun Y, et al. Deep-Learning-Based Drug–Target Interaction Prediction. J Proteome Res [Internet]. 2017 Apr 7;16(4):1401–9. Available from: https://pubs.acs.org/doi/10.1021/acs.jproteome.6b00618.
|
| 41. |
Gawehn E, Hiss JA, Schneider G. Deep Learning in Drug Discovery. Mol Inform [Internet]. 2016 Jan;35(1):3–14. Available from: https://onlinelibrary.wiley.com/doi/10.1002/minf.201501008.
|
| 42. |
Baskin II, Winkler D, Tetko I V. A renaissance of neural networks in drug discovery. Expert Opin Drug Discov [Internet]. 2016 Aug 2;11(8):785–95. Available from: http://www.tandfonline.com/doi/full/10.1080/17460441.2016.1201262.
|
| 43. |
Zeng B, Glicksberg BS, Newbury P, Chekalin E, Xing J, Liu K, et al. OCTAD: an open workspace for virtually screening therapeutics targeting precise cancer patient groups using gene expression features. Nat Protoc [Internet]. 2021 Feb 23;16(2):728–53. Available from: http://www.nature.com/articles/s41596-020-00430-z.
|
| 44. |
Kendall A, Gal Y. What Uncertainties Do We Need in Bayesian Deep Learning for Computer Vision? [Internet]. Cornell University. 2017 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1703.04977v2.
|
| 45. |
Kendall A, Gal Y, Cipolla R. Multi-Task Learning Using Uncertainty to Weigh Losses for Scene Geometry and Semantics [Internet]. Cornell University. 2017 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1705.07115.
|
| 46. |
Szegedy C, Zaremba W, Sutskever I, Bruna J, Erhan D, Goodfellow I, et al. Intriguing properties of neural networks [Internet]. Cornell University. 2013 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1312.6199.
|
| 47. |
Goodfellow IJ, Shlens J, Szegedy C. Explaining and Harnessing Adversarial Examples [Internet]. Cornell University. 2014 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1412.6572.
|
| 48. |
Papernot N, McDaniel P, Sinha A, Wellman M. Towards the Science of Security and Privacy in Machine Learning [Internet]. Cornell University. 2016 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1611.03814v1.
|
| 49. |
Xu W, Evans D, Qi Y. Feature Squeezing: Detecting Adversarial Examples in Deep Neural Networks [Internet]. Cornell University. 2017 [cited 2022 Aug 26]. Available from: https://arxiv.org/abs/1704.01155.
|
| 50. |
Phillips A, Borry P, Shabani M. Research ethics review for the use of anonymized samples and data: A systematic review of normative documents. Account Res [Internet]. 2017 Nov 17;24(8):483–96. Available from: https://www.tandfonline.com/doi/full/10.1080/08989621.2017.1396896.
|